Forschung, die weiterbringt
Pro Jahr arbeiten mehr als 400 Forscher*innen gemeinsamen mit Kooperationspartnern aus Wirtschaft und Gesellschaft an über 500 laufenden Forschungprojekten.
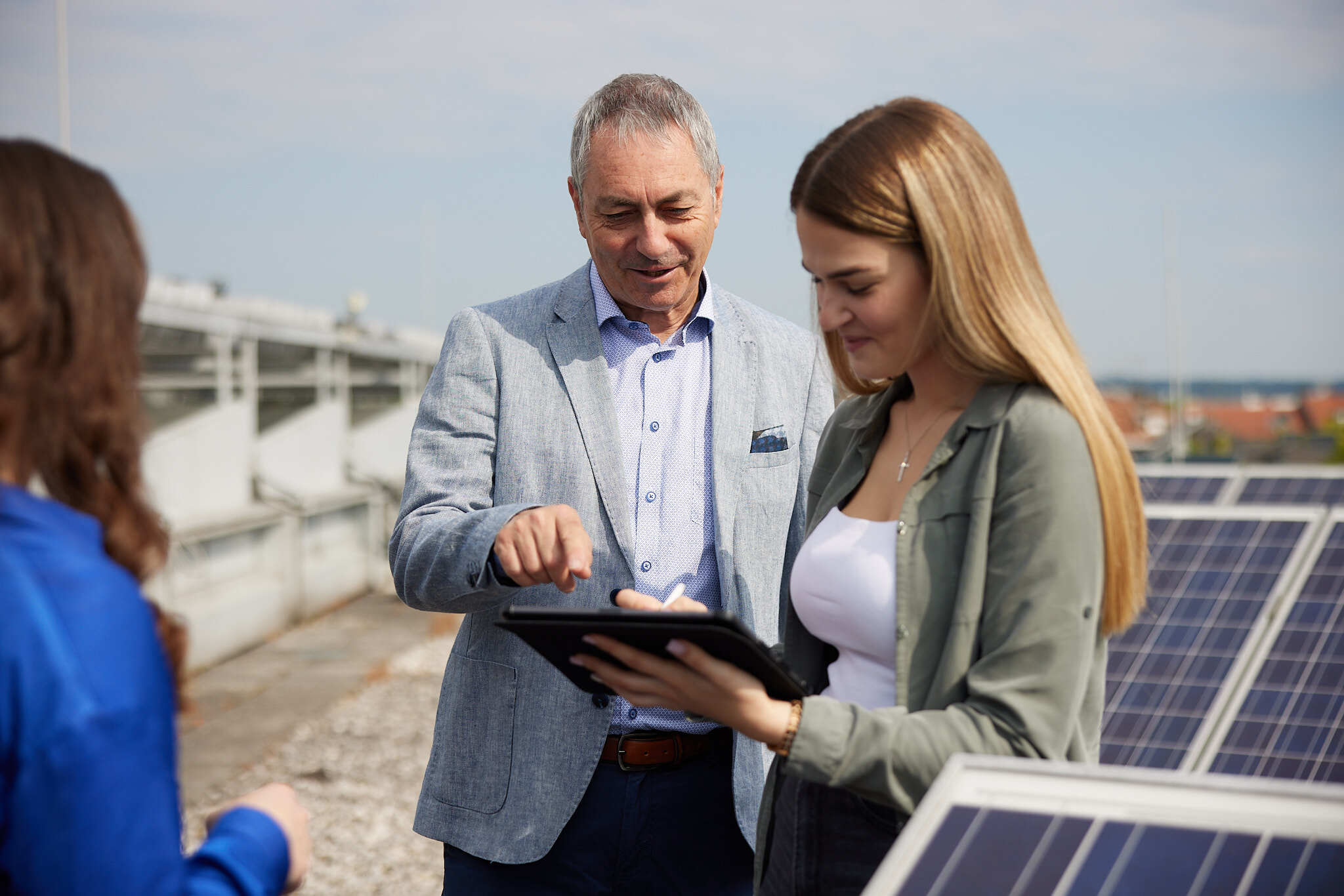

Wir gestalten die Zukunft aktiv mit
Nachhaltigkeit, Digitalisierung, Gesundheit oder soziale Gerechtigkeit. Unser Ziel ist es, konkrete Herausforderungen unserer Zeit zu lösen und die wissenschaftliche Erkenntnisse unmittelbar für Wirtschaft und Gesellschaft zugänglich und nutzbar zu machen.
Dazu bearbeiten wir Problem- und Fragestellungen in verschiedensten Dimensionen und Rahmenbedingungen: von Einzelprojekten für Unternehmen über mehrjährige, geförderte Forschungsprojekte bis zu dauerhaften F&E-Kooperationen in Forschungszentren, die an unseren Fakultäten angesiedelt sind. Auf diese Weise können innovative Produkte, Dienstleistungen oder Prozesse entwickelt werden. Durch enge Beziehungen zur regionalen Wirtschaft und lokalen Gemeinschaften unterstützen wir verstärkt den Transfer zwischen Hochschule und Industrie.
Mit ihrer Zugkraft im Bereich F&E zählt die FH OÖ seit Jahren zu den forschungsstärksten Fachhochschulen im deutschsprachigen Raum.
... international und interdisziplinär
Wir verknüpfen verschiedenste Forschungsbereiche aus Wirtschaft, Technik und Sozialem und bauen gleichzeitig auf Partnerschaften und Kooperationen mit internationalen Hochschulen oder Unternehmen. Neben der Arbeit an internationalen Projekten ergibt sich für unsere Mitarbeiter*innen auch die Möglichkeit von Forschungsaufenthalten an ausländischen Hochschulen. Das so gewonnene, vielseitige Know-how fließt wiederum direkt in die Umsetung innovativer Lösungen, die noch näher an der Praxis und damit an den Bedürfnissen der Anwender*innen stehen. Und letztendlich auch den entscheidenden Vorteil für Wirtschaft und Gesellschaft bringen.
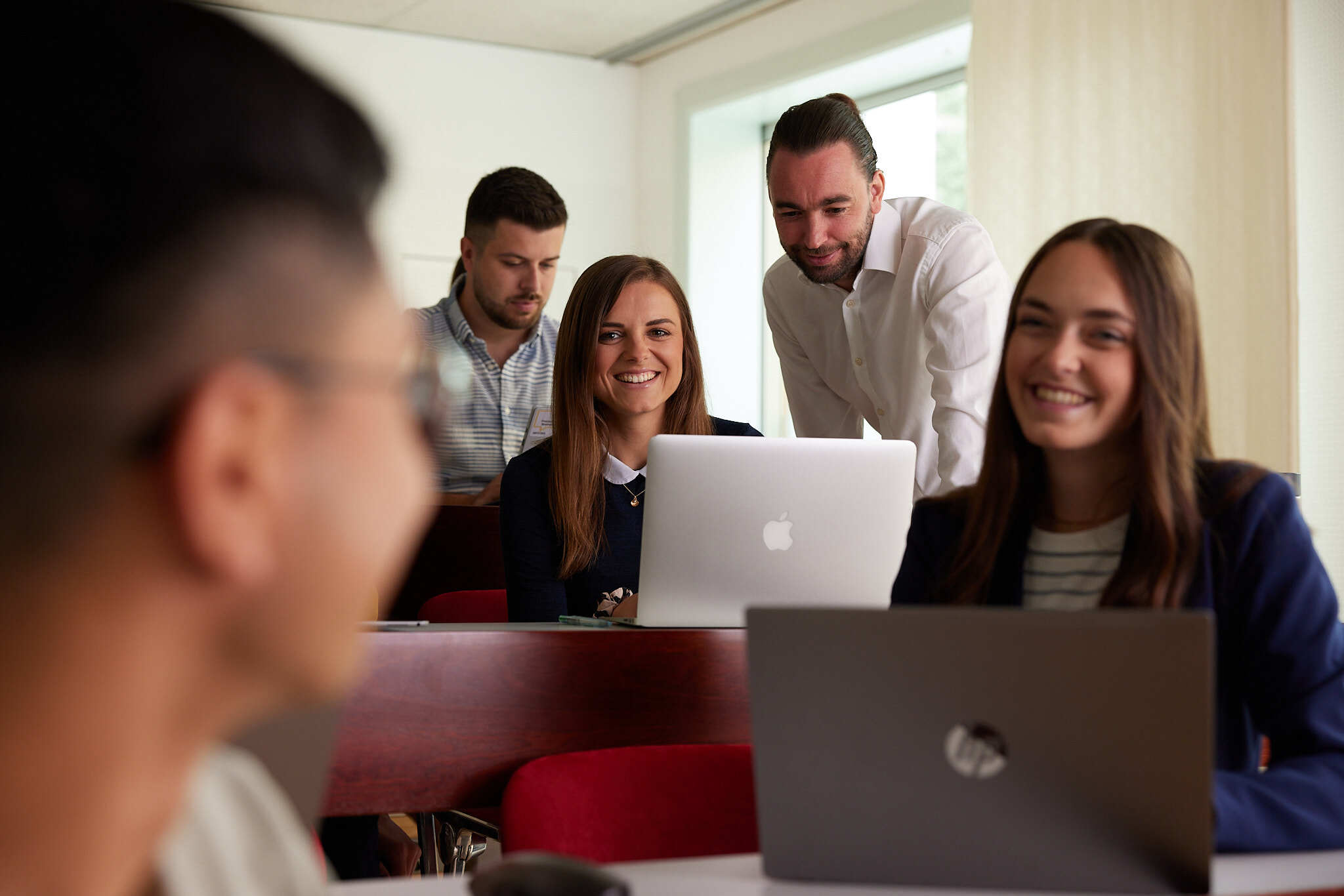

... ein Sprungbrett für junge Talente
Studierende der FH OÖ haben bereits während ihrer Ausbildung die Möglichkeit im Bereich Forschung & Entwicklung mitzuarbeiten und die gelernte Theorie in die Praxis umzusetzen. Ob in Form eines Projekts, Praktikums oder einer Abschlussarbeit: Angewandte Forschung gibt einen wertvollen Einblick in die Berufsmöglichkeiten und ermöglicht unseren Studierenden sich mit Fachleuten, Professor*innen und anderen Nachwuchsforscher*innen zu vernetzen.
Zur Verknüpfung von Lehre und Forschung wurde an jeder der vier Fakultät ein eigenes Research Center mit spezifischen Forschungsschwerpunkten errichtet.